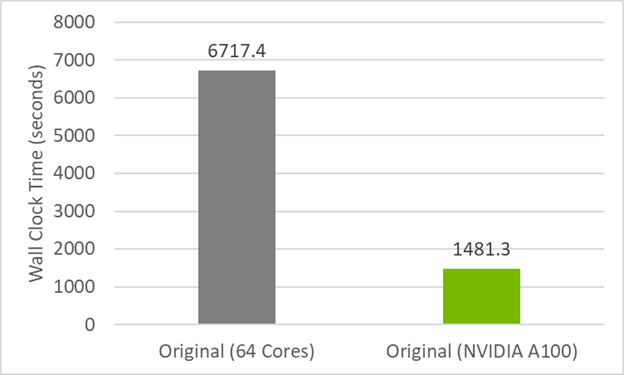
With JAX, Veros reaches Fortran speed for both few and many CPU cores,... | Download Scientific Diagram

Distribution of batch CPU time.P ercentages in child nodes indicate the... | Download Scientific Diagram

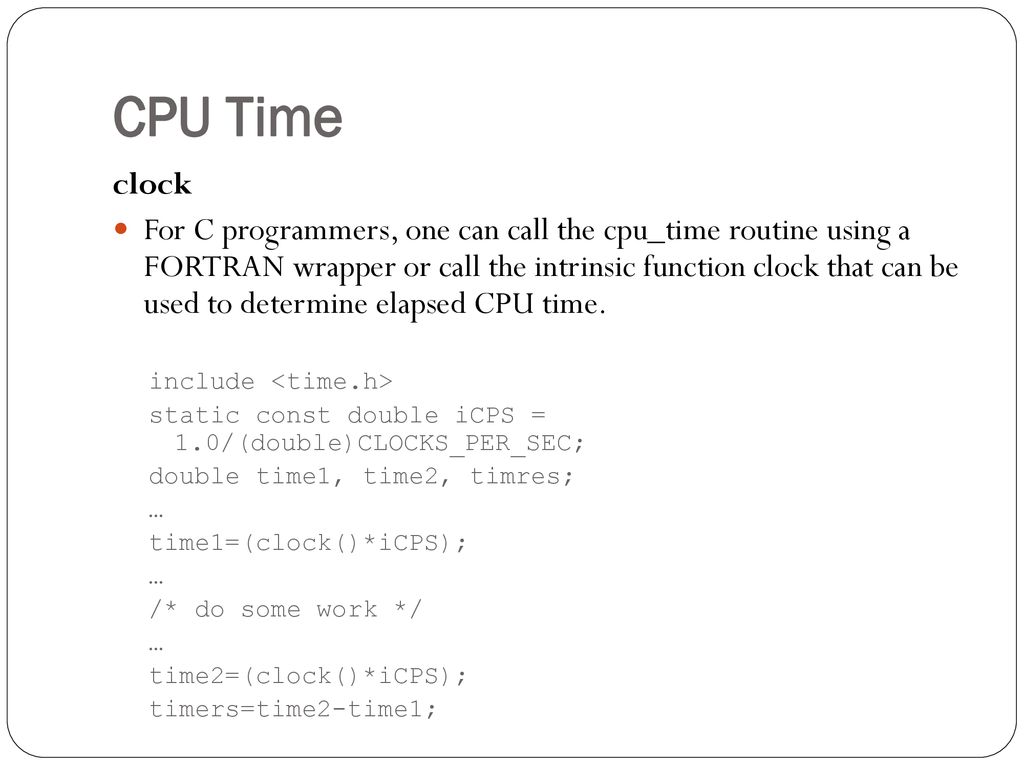
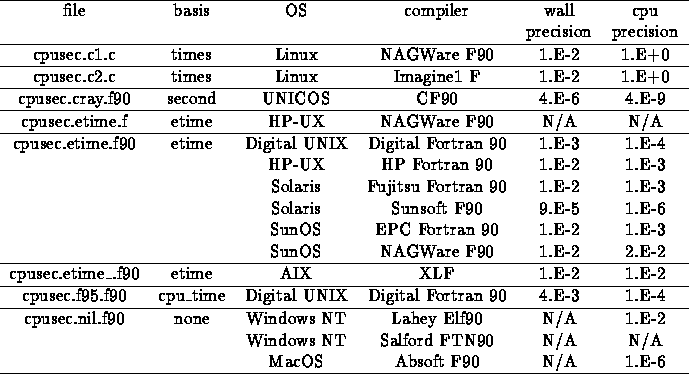
CPU times measured with the Fortran intrinsic cpu time and normalized... | Download Scientific Diagram

performance - Huge slowdown when packing arguments into derived type with pointers in Fortran - Stack Overflow

Table 1 from OpenMP GNU and Intel Fortran programs for solving the time-dependent Gross-Pitaevskii equation | Semantic Scholar
![PDF] Vectorized link cell Fortran code for molecular dynamics simulations for a large number of particles | Semantic Scholar PDF] Vectorized link cell Fortran code for molecular dynamics simulations for a large number of particles | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/5feebb879ddac769e2a26af58e07437a22ff395b/10-Table1-1.png)
PDF] Vectorized link cell Fortran code for molecular dynamics simulations for a large number of particles | Semantic Scholar

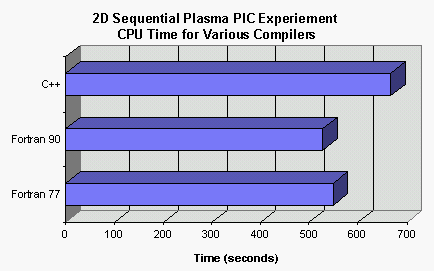
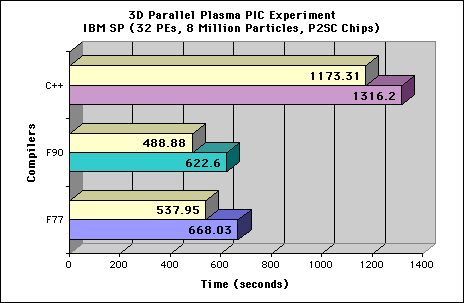
Ratio of the CPU time of each case to that of the "inlinc Fortran 77"... | Download Scientific Diagram

Table 2 from Vector pipelining, chaining, and speed on the IBM 3090 and Cray X-MP | Semantic Scholar

CPU-time reported for the CCSM simulations, compiled with the Intel... | Download Scientific Diagram

Ratio of the CPU time of each case to that of the "inlinc Fortran 77"... | Download Scientific Diagram
![PDF] Vectorized link cell Fortran code for molecular dynamics simulations for a large number of particles | Semantic Scholar PDF] Vectorized link cell Fortran code for molecular dynamics simulations for a large number of particles | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/5feebb879ddac769e2a26af58e07437a22ff395b/11-Table2-1.png)
PDF] Vectorized link cell Fortran code for molecular dynamics simulations for a large number of particles | Semantic Scholar

Benchmarking a simple PDE algorithm in Julia, Python, Matlab, C++, and Fortran - #18 by ChrisRackauckas - Numerics - Julia Programming Language