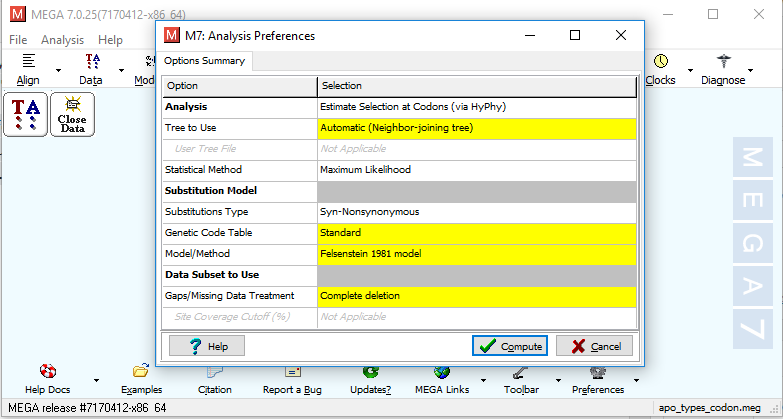
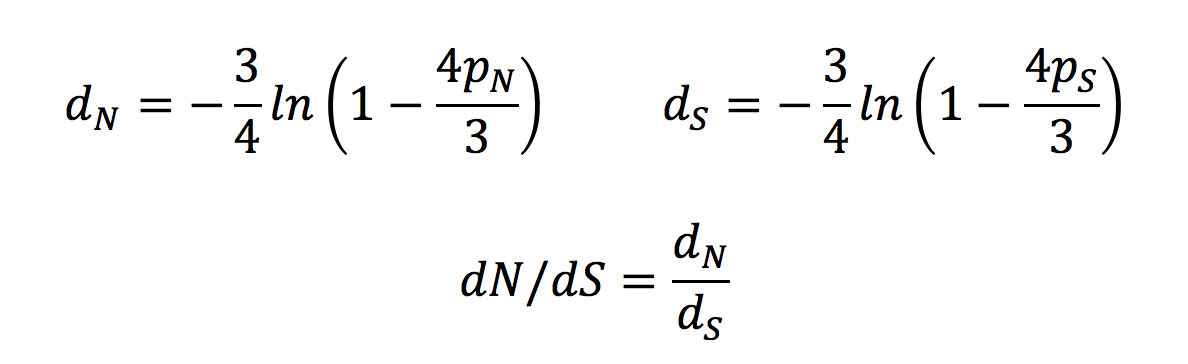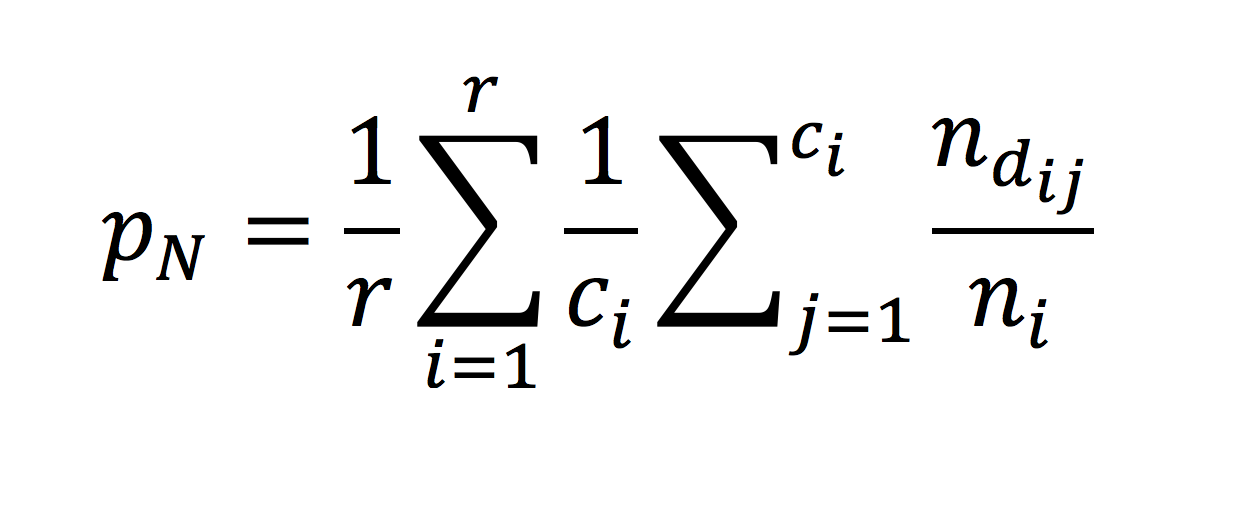A comparison of one-rate and two-rate inference frameworks for site-specific dN/dS estimation | bioRxiv
GitHub - adelq/dnds: Calculate dN/dS ratio precisely (Ka/Ks) using a codon-by-codon counting method.
1 Any questions? Analysis of variation in the dn/ds ratio between sites or lineages dN/dS ratio (ω) estimated across all sites
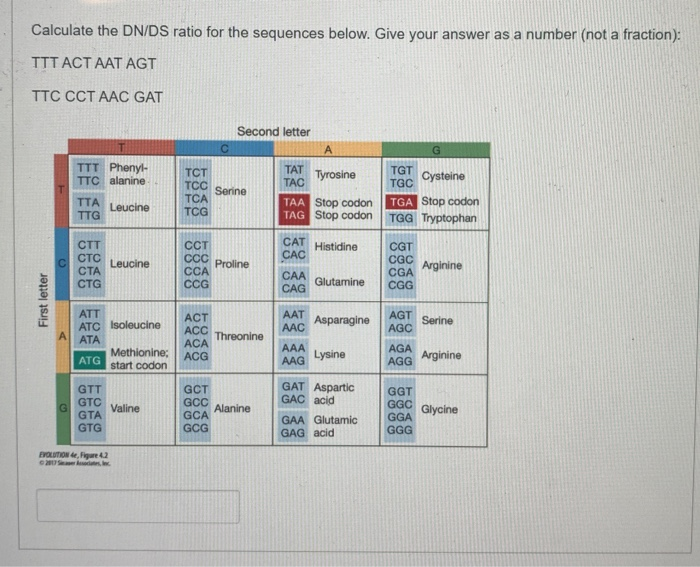
![University Human Biology: dN/dS rate] Can someone explain to me how these calculations are correct? Are non-synonymous changes when there is a change in the amino acid? How are some of these University Human Biology: dN/dS rate] Can someone explain to me how these calculations are correct? Are non-synonymous changes when there is a change in the amino acid? How are some of these](https://preview.redd.it/24uzas5dti231.png?auto=webp&s=0318e5dc55044bbeab123a08969026bb035b7fdd)